England’s rivers are under pressure. Pollution from agricultural runoff, treated and untreated wastewater, and a changing climate are testing the resilience of freshwater ecosystems that millions of species and people depend on. Chemical contaminants can enter rivers in short pulses following rainfall or arise more persistently from diffuse sources across entire catchments, making their ecological impacts difficult to detect and monitor at scale. Regulatory frameworks for assessing river health rely primarily on surveys of macroinvertebrates, diatoms, and macrophytes. These tools are well established but are not always sensitive to early or diffuse ecological stress.
Biofilms, the greenish layer that grows on the surfaces of rocks in rivers and streams, are home to complex assemblages of microbes that are central to the ecological functioning of rivers. Bacteria within these biofilms drive nutrient cycling, process contaminants, and form the base of aquatic food webs. These communities are highly sensitive to their environment, responding rapidly to a wide range of pressures, including changes in nutrients, pollutants, temperature, and hydrology. Despite their ecological importance, they remain poorly characterised and are rarely incorporated into routine monitoring.
Advances in environmental DNA sequencing are beginning to change this. It is now possible to recover thousands of microbial genomes from a single sample of water, sediment, or biofilm, allowing us to identify which microbes are present and the genes they carry. This means we can begin to understand not just which microbes are there, but what they are doing, providing a powerful approach to assessing ecosystem health and detecting subtle, early ecological changes.
The NERC PACIFIC Project
The PACIFIC (PAthways of Chemicals Into Freshwaters and their ecological ImpaCts) project investigates how human‑made chemicals enter freshwater environments and affect microbial communities that underpin ecosystem health and biogeochemical cycling. Funded by UKRI Natural Environment Research Council (NERC) as part of the Understanding Changes in Quality of UK Freshwaters programme, PACIFIC combines large‑scale catchment monitoring with advanced chemical analysis and molecular techniques to link chemical sources, pathways, and exposure to ecological responses in rivers. By focusing on freshwater microbes, an often overlooked but functionally critical component of ecosystems, the project aims to improve understanding of how complex chemical mixtures influence microbial structure and function, providing evidence to inform future monitoring approaches, risk assessment, and freshwater management.
The national survey
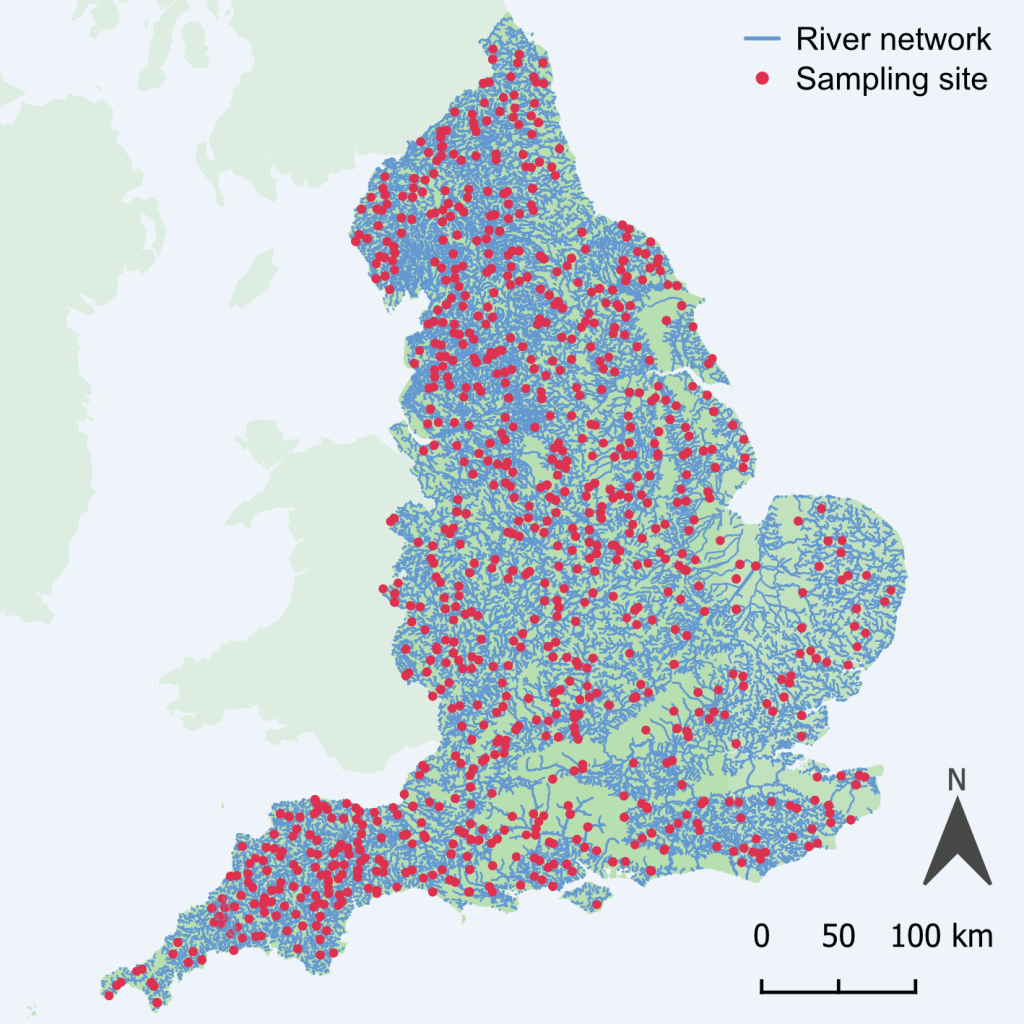
As part of PACIFIC, the UK Centre for Ecology & Hydrology (UKCEH) and the Environment Agency (EA) worked in partnership to analyse thousands of biofilms collected from across the EA’s River Surveillance Network (Figures 1 and 2), an initiative designed to monitor long-term changes in river health.
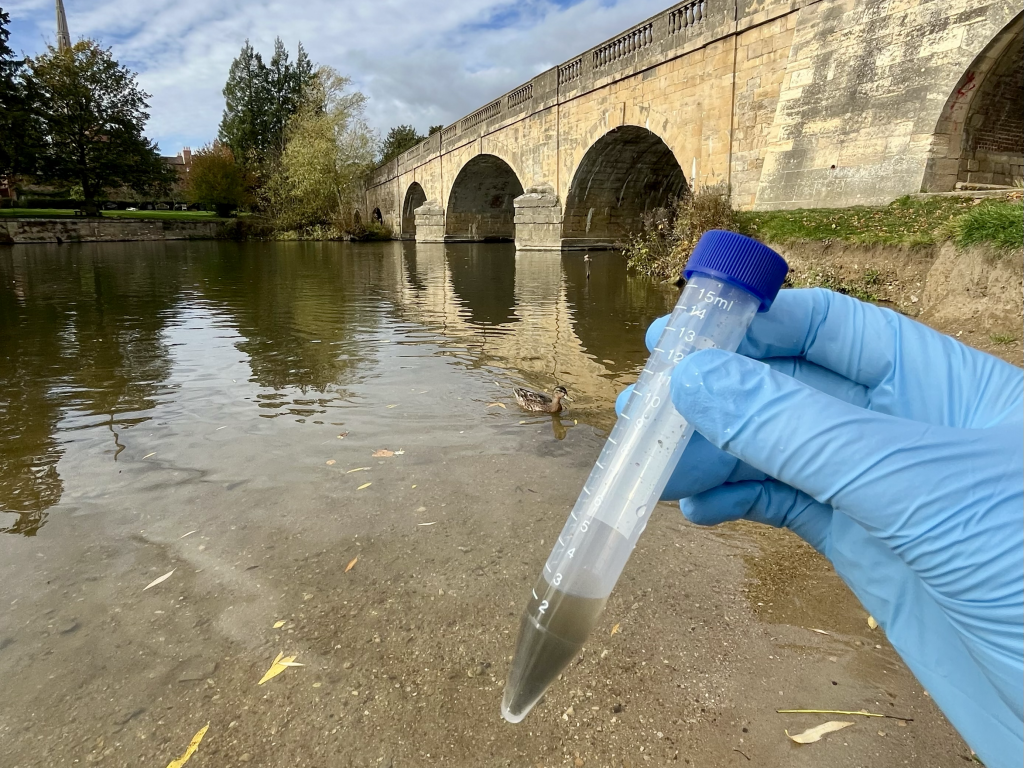

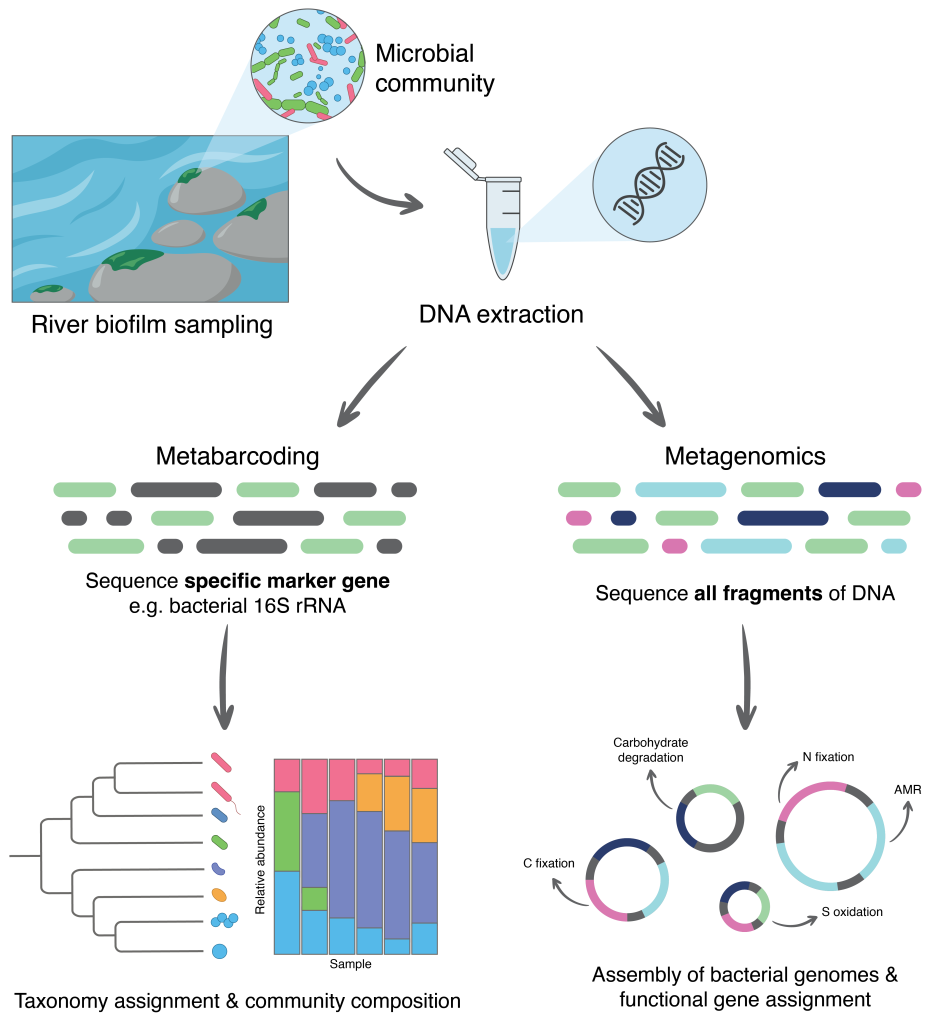
The research characterised the diversity and function of river biofilm bacteria at a national scale using a combination of two high-throughput DNA sequencing approaches. Metabarcoding targets a specific marker gene to identify which microbes are present in a sample, and metagenomic sequencing captures all DNA extracted from a sample and enables the assembly of individual bacterial genomes, revealing not only which microbes are present, but also the genes they encode (Figure 3). These genes provide insight into the functions the microbes may perform in the environment. Combined with water chemistry and catchment land use data, this approach offers a detailed picture of the ecology of river biofilm bacteria, and a means of understanding the condition of the rivers they inhabit.
Diversity and function of river biofilm bacteria
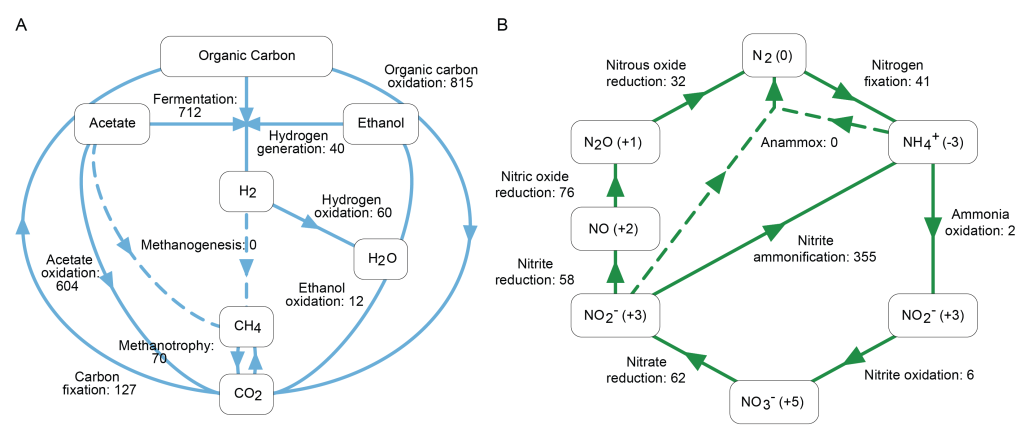
The national survey revealed a remarkable diversity of bacteria, including many not previously described. These communities carry a wide range of genes associated with carbon and nitrogen cycling (Figure 4) and breakdown of organic matter, highlighting their role in making nutrients available to higher trophic levels such as grazing micro and macroinvertebrates, including insect larvae, and the fish that feed on them.
Genes linked to antimicrobial resistance and the uptake, transformation, and breakdown of contaminants were widespread. This suggests that biofilm bacteria can influence the persistence and mobility of contaminants in ways that may either mitigate or exacerbate their impacts, with implications for both water quality and the spread of antimicrobial resistance in river systems.
The biogeographic distribution of biofilm bacteria across the river network was strongly shaped by the surrounding environment. Water chemistry, nutrient concentrations, wastewater inputs, and catchment land cover, whether agricultural or urban, all emerged as significant drivers of which bacteria were present and in what abundance. Because these bacterial communities both respond to and regulate key ecosystem processes, changes in their composition are not only indicators of environmental pressures but may also signal shifts in how nutrients and contaminants are processed in rivers.
Biofilms as indicators of river health
Biofilms, attached to submerged surfaces, allow microbes to persist in the river longer than those suspended in the water column. This persistence means that biofilm communities respond to environmental signals accumulated over space and time, reflecting both local conditions and inputs from the catchment. As a result, they provide a highly informative and temporally stable indicator of environmental change.
Within these communities, candidate indicator taxa were identified that show strong, directional responses to environmental pressures such as increasing nitrate concentrations. Clear change points – where responses were most pronounced – emerged at specific points along the nitrate concentration gradient. The change points represent thresholds where relatively small changes in environmental conditions are associated with marked and detectable shifts in biofilm communities.
These findings have important implications for river monitoring. The presence of responsive bacterial taxa and identifiable thresholds suggests that biofilm communities could be used not only to detect change, but to determine when ecosystems are approaching critical transitions. Given the central role of these microbes in nutrient cycling and contaminant processing, such shifts may precede and help explain broader ecological impacts. This creates the potential for earlier and more sensitive warning signals of deterioration, enabling timely and targeted management interventions (Figure 5).
Translating this research into regulatory practice will require further validation, standardisation, and engagement with monitoring agencies and policymakers. Nonetheless, this work represents an important step toward integrating microbial indicators into freshwater assessment frameworks, with the potential to strengthen the detection, interpretation, and management of environmental change.

This work was supported by the PACIFIC project, which was funded by the Natural Environment Research Council, under grant numbers NE/X015947/1, NE/X015890/1, NE/X015777/1 and NE/X015874/1, and the Environment Agency Science Project SC220034.
Header image: © Alex Cooper | Adobe Stock

